Ryan Hisner and others recently observed something unusual in the GISAID sequence data: the emerging SARS-CoV-2 lineage BA.3.2 appeared to have a disproportionate number of sequences from children compared to other circulating lineages. We decided to take a closer look.
Analyzing the Data
To investigate, we queried GISAID for all countries that included patient age metadata in their sequence submissions and identified the five countries with the most BA.3.2 sequences: Luxembourg, the Netherlands, Ireland, France, and the USA. For each country, we compared the age distribution of BA.3.2 sequences to non-BA.3.2 sequences from the same country and time period (December 2025 to present).
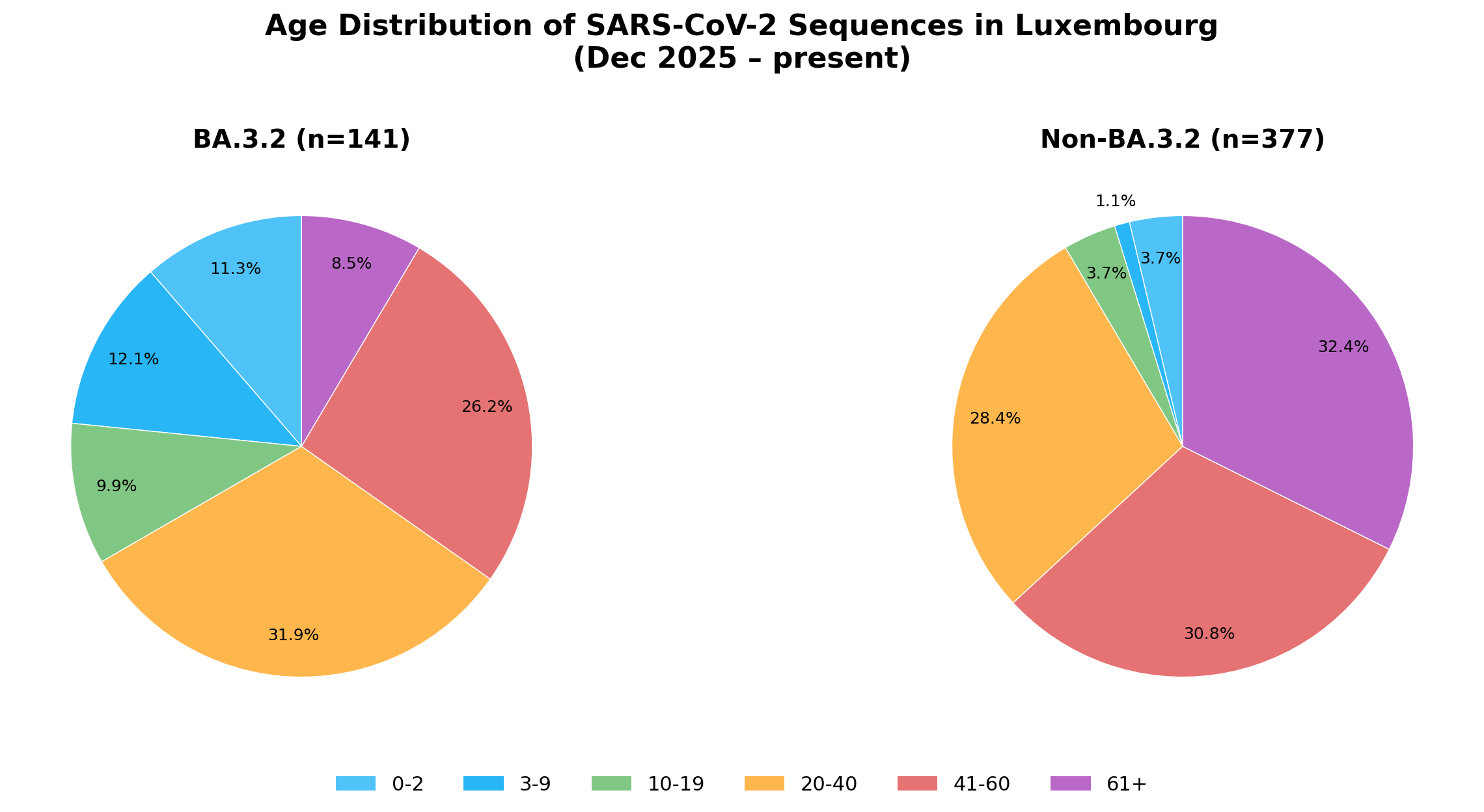
The results were striking. In Luxembourg, children under 10 made up 23.4% of BA.3.2 sequences (n=141) versus just 4.8% of non-BA.3.2 (n=377), an over 4-fold increase.
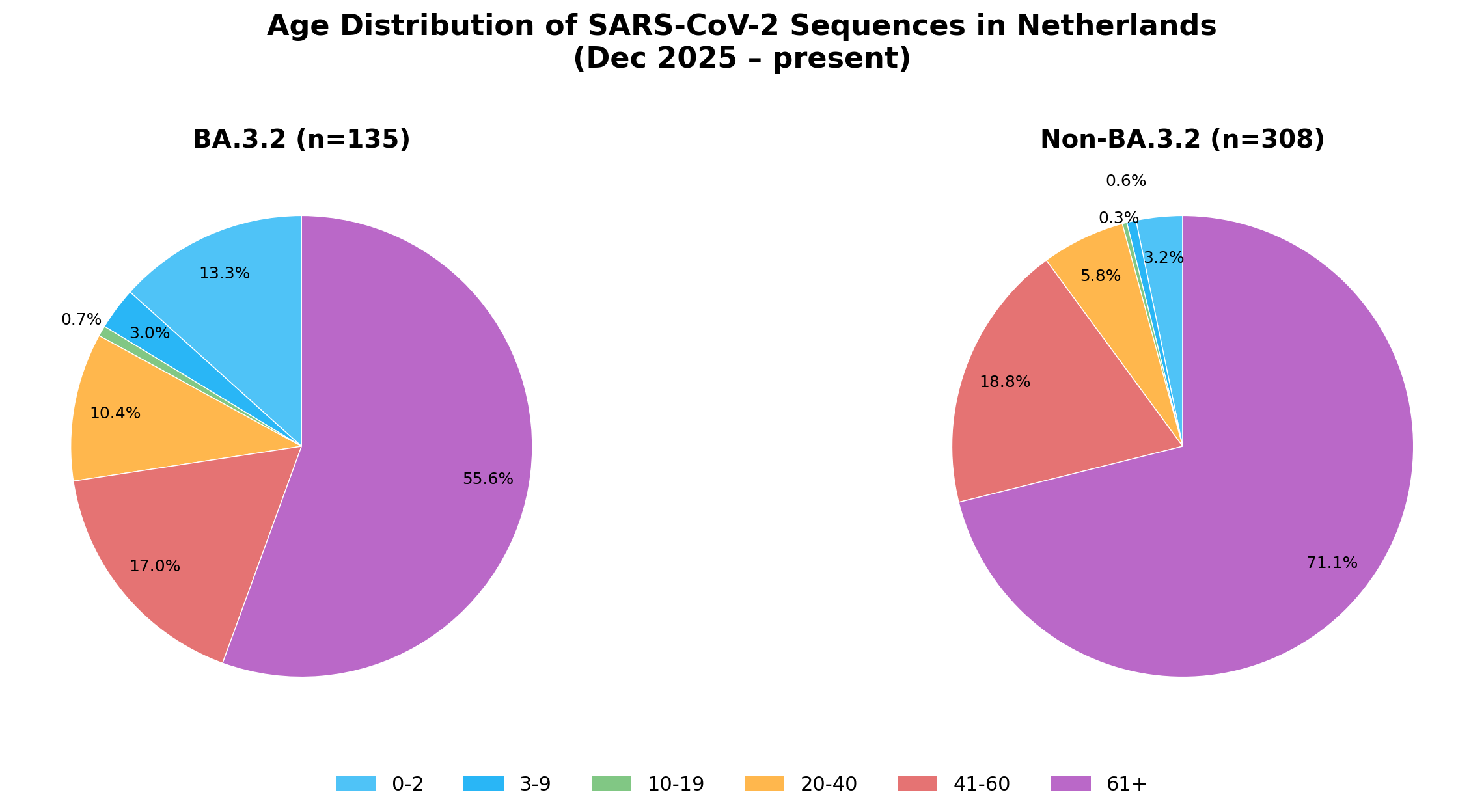
In the Netherlands, 16.3% of BA.3.2 sequences (n=135) came from children under 10, compared to just 3.8% of non-BA.3.2 (n=308), also a 4-fold increase.
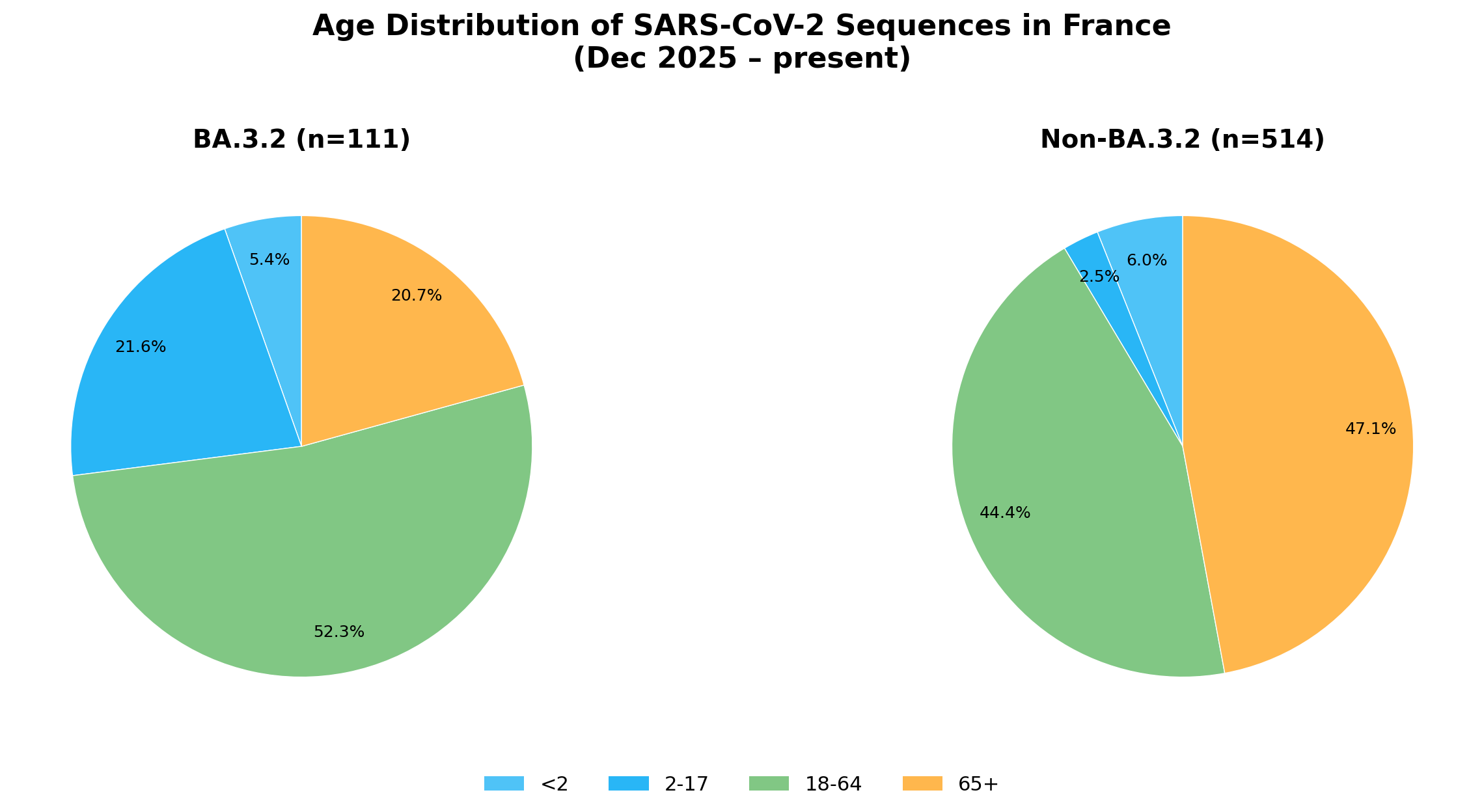
France showed a clear skew in the pediatric age group: 27.0% of BA.3.2 sequences (n=111) were from patients under 18, versus just 8.5% of non-BA.3.2 (n=514). The difference was most pronounced in children 2-17, which was 21.6% of BA.3.2 sequences and only 2.5% of non-BA.3.2 sequences, an 8-fold increase in this category.
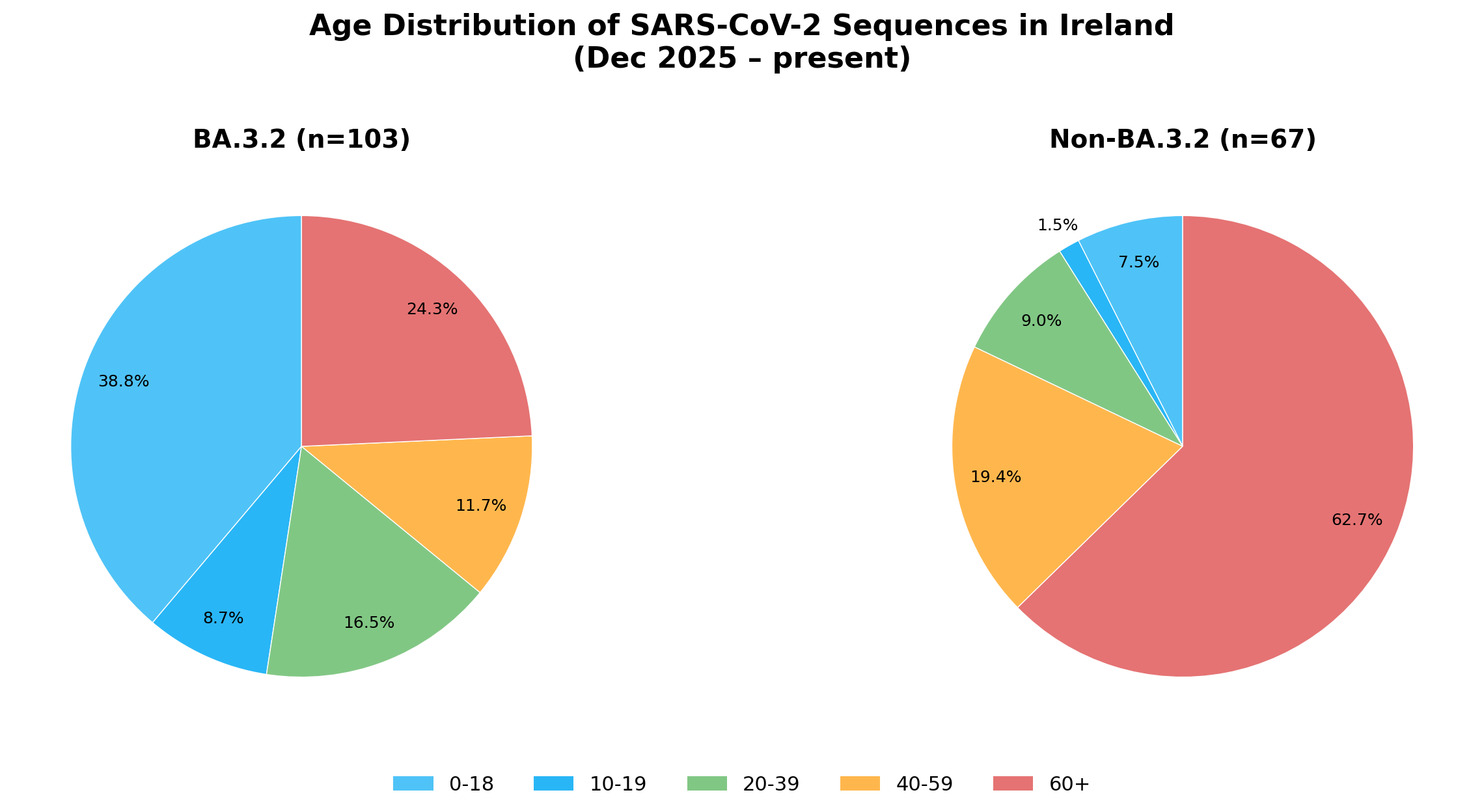
In Ireland, 38.8% of BA.3.2 sequences (n=103) were from children aged 0-18, compared to just 7.5% of non-BA.3.2 (n=67), a 5-fold increase.
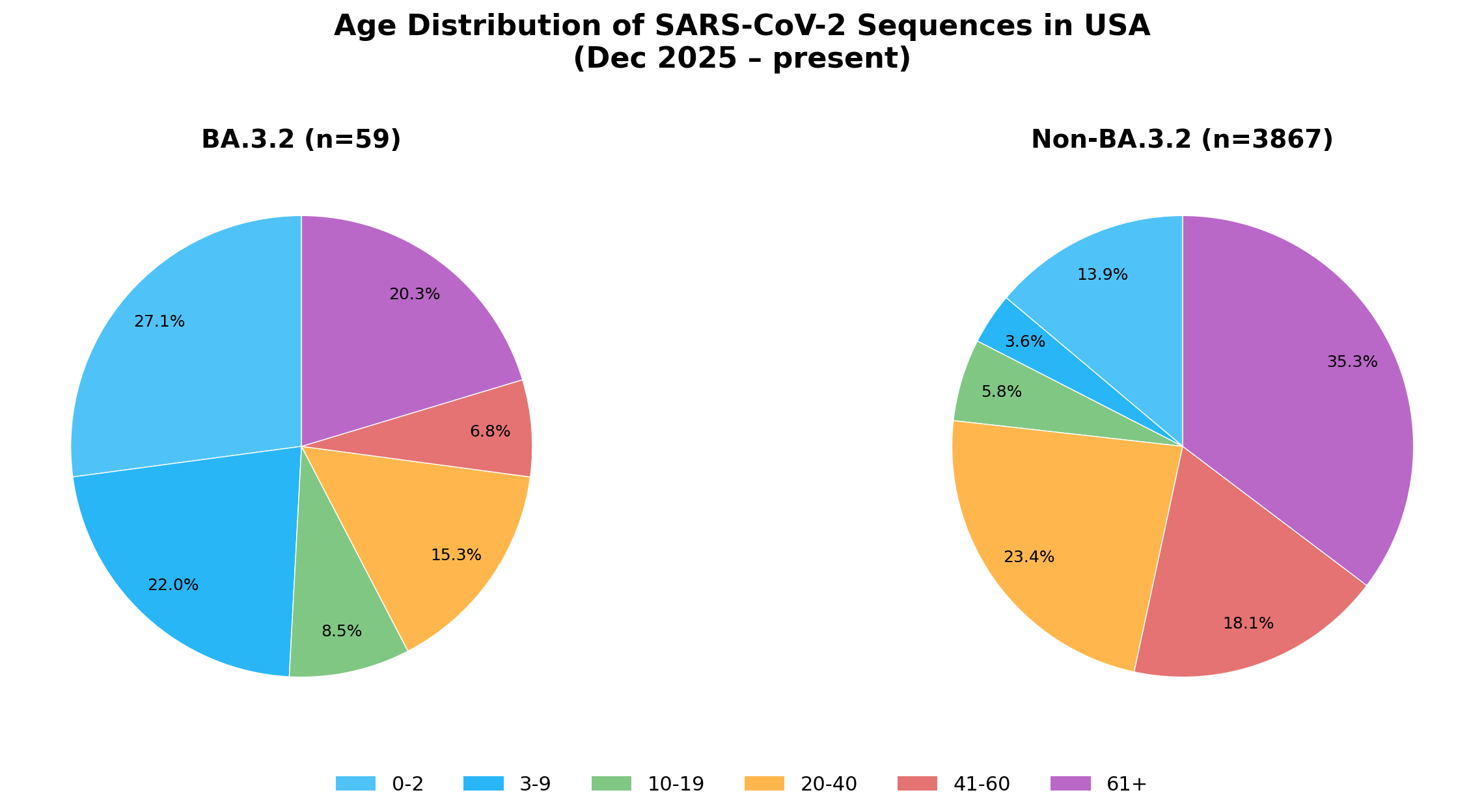
In the USA, 49.1% of BA.3.2 sequences (n=59) were from children under 10, compared to 17.5% of non-BA.3.2 sequences (n=3,867). The most pronounced group was 3-9, which was 22% of BA.3.2 sequences but only 3.6% of non-BA.3.2 sequences, a 6-fold increase.
Why Might This Be Happening?
Hypothesis 1: Immunological Naivety
One straightforward explanation is that younger individuals have had fewer prior SARS-CoV-2 infections and therefore less immune experience. If BA.3.2 is antigenically distinct enough from previously circulating lineages, children — who may have only been exposed to one or two prior lineages — could be more susceptible to infection by something new, leading to higher rates of clinical presentation and sequencing.
To test this, we looked back at the last major lineage shift: the displacement of XBB lineages by BA.2.86/JN.1 in late 2023 to early 2024. If immunological naivety were the primary driver, we would expect to see a similar skew toward younger age groups in BA.2.86 sequences compared to XBB.
We did not. The age distributions of XBB (n=4,772) and BA.2.86 (n=8,476) were virtually identical, with no enrichment of children in the newer lineage.
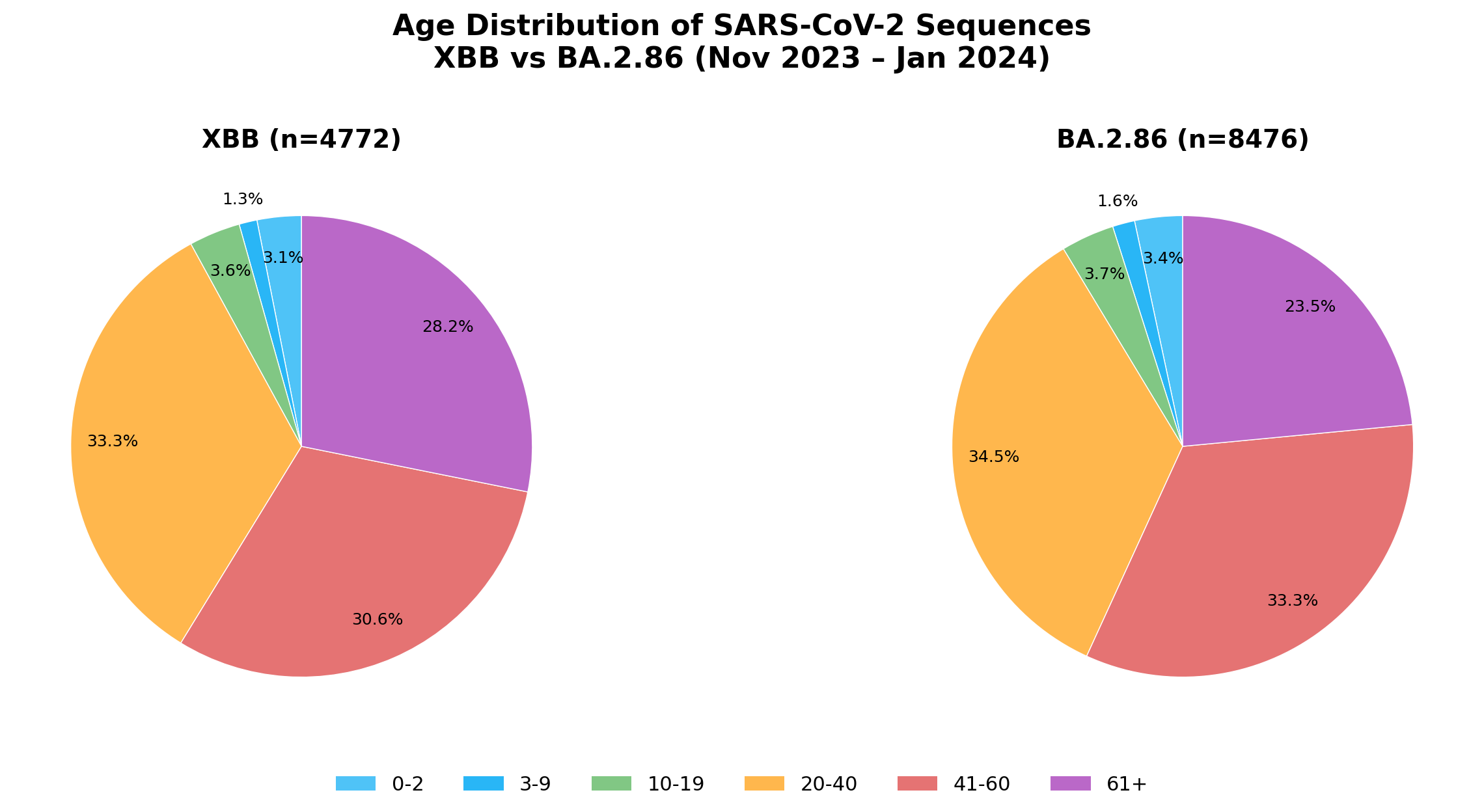
This suggests that immunological naivety alone does not explain the BA.3.2 age skew.
Hypothesis 2: The ORF7/8 Deletion
Ryan Hisner has proposed that the ORF7/8 deletion carried by BA.3.2 may be responsible for the age-skewed pattern. To explore this, we looked for a historical comparator: GW.5.1.1, another lineage that carried a similar ORF7/8 deletion. We compared its age distribution to HV.1.1, a lineage that circulated at the same time but lacked this deletion.
The results were suggestive. Among US sequences, GW.5.1.1 (n=35) showed a notably younger age distribution than HV.1.1 (n=468), with a higher proportion of sequences from children and young adults.
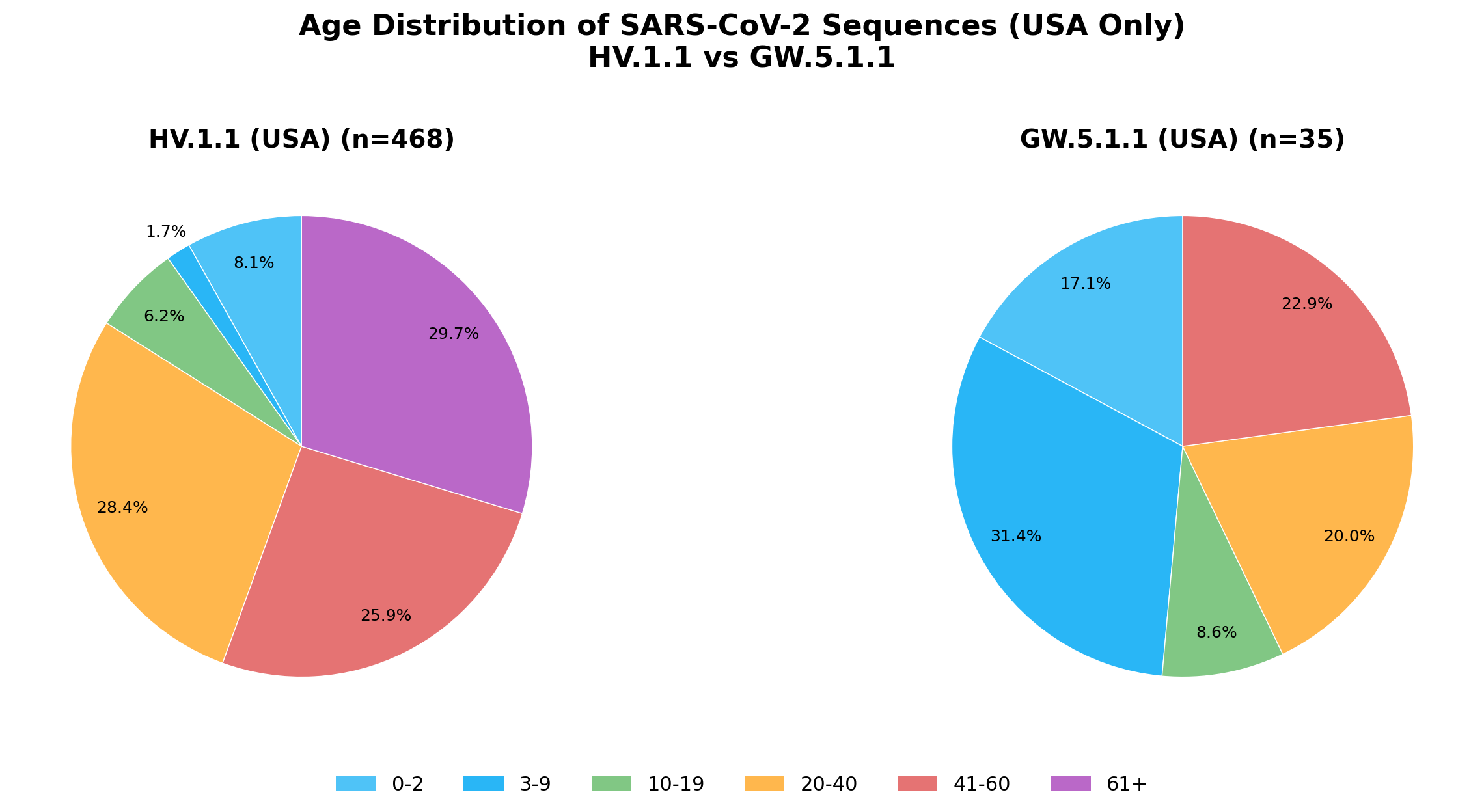
Conclusions
The data clearly show that BA.3.2 sequences are enriched for younger individuals compared to other lineages circulating at the same time and place. The consistency of this pattern across five countries with different sequencing practices and healthcare systems makes it unlikely to be an artifact of sampling bias.
The immunological naivety hypothesis — that children are simply more susceptible to any new lineage — is not supported by the XBB/BA.2.86 comparison, where no age skew was observed during a comparable lineage transition.
The ORF7/8 deletion hypothesis, proposed by Ryan Hisner, is more consistent with the data. The similar age skew seen in GW.5.1.1, another lineage with a comparable deletion, suggests that this genomic feature may play a role in altering the age distribution of infections. Further investigation into how ORF7/8 deletions affect viral tropism, immune evasion, or disease presentation across age groups is warranted.
Data & Acknowledgments
Data were downloaded from GISAID’s EpiCoV database. We gratefully acknowledge all data contributors, i.e., the Authors and their Originating laboratories responsible for obtaining the specimens, and their Submitting laboratories for generating the genetic sequence and metadata and sharing via the GISAID Initiative, on which this research is based.
All genome sequences and associated metadata supporting the findings of this study can be accessed through the persistent digital object identifier https://doi.org/10.55876/gis8.260406wc. GISAID also communicates the aggregation of GISAID accession numbers (EPI_ISL_IDs) through the corresponding EPI_SET_260406wc identifier to facilitate both the acknowledgment of all data contributors and the direct retrieval of the underlying data from GISAID used in this study.